In my last post I talked about Chemical Shift Selective Filter (CSSF) experiments1 and their ability to excite selectively signals that are overlapped with other signals in the spectrum. When combined with mixing schemes such as NOESY/TOCSY, this high degree of selectivity means that CSSF experiments can extract important structural information from the spectrum that is not obtainable using conventional selective excitation methods. Despite their great power, though, CSSF experiments have a number of drawbacks that have limited their use. The principal issues are the long measurement times needed to measure CSSF spectra – CSSF requires multiple spectra to be collected and co-added in order to achieve the desired selectivity – and some complexities around experiment optimization and data processing.
Fortunately, a new approach to “ultra-selective” excitation has recently been published that circumvents many of the limitations of CSSF. The Gradient-Enhanced Multiplet-Selective Targeted-Observation NMR Experiment (GEMSTONE)2,3 provides comparable selectivity to CSSF but, unlike CSSF, that selectivity is achieved in a single scan. This makes GEMSTONE much quicker to run than CSSF, with the additional benefit that no special processing is required for GEMSTONE data.
GEMSTONE
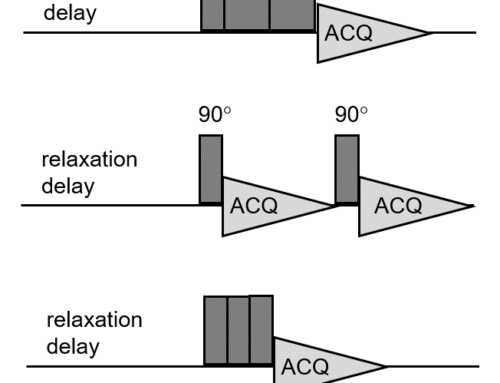
The basic GEMSTONE pulse sequence is shown in Figure 1 .

Figure 1. GEMSTONE pulse sequence
I won’t go into exact details of how GEMSTONE works and interested readers should check out the original papers2,3 on GEMSTONE, but the basic idea is that the combination of the frequency-swept pulses (indicated with diagonal arrows in Figure 1) and the weak field gradients (G1) means that, at the start of the acquisition period, spins at different positions along the direction of the gradient axis will have experienced different t1 evolution periods. This can be compared directly with the CSSF approach of performing separate experiments with incremented t1 delays. As with CSSF, signals experience a modulation of their phase that depends on both how far away from the transmitter they are and how long the effective t1 period is. However, unlike CSSF, where the suppression of the off-resonance signals relies on co-adding the data from separate experiments, in GEMSTONE the different increments are spatially encoded within each scan and so the suppression occurs within a single scan!
GEMSTONE NOE
The power of GEMSTONE becomes apparent when it’s combined with mixing schemes such as TOCSY or NOESY to create both fast and ultra-selective versions of those experiments. Figure 2 shows the pulse sequence for the GEMSTONE NOE experiment.

Figure 2. GEMSTONE NOE pulse sequence
GEMSTONE NOE Example
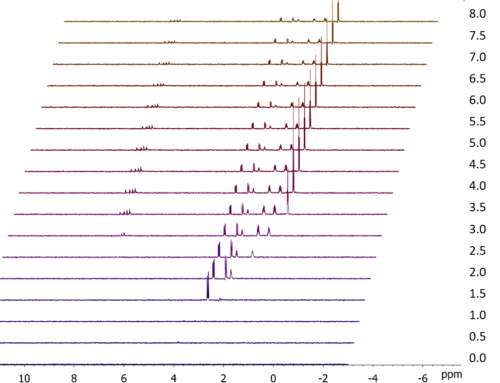
Figure 3 shows a comparison of a conventional selective NOESY spectrum with a set of three different GEMSTONE NOE spectra collected on a sample of 17β-estradiol. The data were collected on a JEOL ECZ500R spectrometer equipped with a ROYAL HFX probe and Delta software and processed using JASON software. The GEMSTONE NOE spectra were each collected using 128 scans with an experiment time of 14 minutes, while the standard 1D NOESY experiment was collected using 32 scans in 4 minutes. These measurement times compare with the one hour needed to collect the CSSF-NOESY spectrum shown here.
The three signals around 1.8 ppm are sufficiently overlapped to make individual excitation of each impossible using the conventional method. However, GEMSTONE is able to disentangle this overlapped region and excite each signal separately, thereby allowing NOE correlations between each to be unambiguously observed.

Figure 3. Comparison of conventional selective NOESY with GEMSTONE NOE. The conventional experiment is unable to excite separately the three overlapped peaks at around 1.8 ppm, whereas the GEMSTONE NOE experiment allows separate excitation of each peak.
To find out more about GEMSTONE experiments, or to obtain copies of the GEMSTONE pulse programs for JEOL ECZ spectrometers, contact us at the usual address!
References
- T. Robinson, T. N. Pham, D. Uhrin, J. Magn. Reson., 2004, 170, 97-103.
- Kiraly, N. Kern, M. P. Plesniak, M. Nilsson, D. J. Procter, G. A. Morris and R. W. Adams, Angew. Chem. Int. Ed. 2021, 60, 666–669.
- Kiraly, M. Nilsson, G. A. Morris and R. W. Adams, Chem. Commun. 2021, 57, 2368-2371.